Turnover statistics
Average time to find at least 2 reviewers after submission = 26 days (median = 17)
Average time from submission to 1st decision = 68 days (median = 57)
Latest recommendations

| Id | Title | Authors▲ | Abstract | Picture | Thematic fields | Recommender | Reviewers | Submission date | |
|---|---|---|---|---|---|---|---|---|---|
09 May 2023
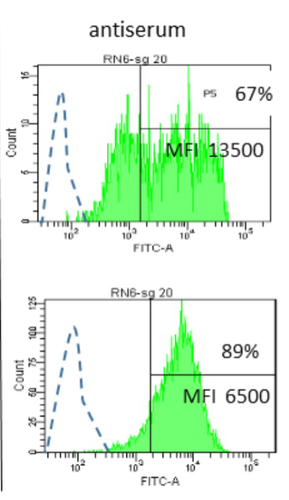
Interactions between Mycoplasma mycoides subsp. mycoides and bovine macrophages under physiological conditionsPhilippe Totté, Tiffany Bonnefois, Lucia Manso-Silvan https://doi.org/10.1101/2022.12.06.519279Interaction of bovine macrophages with Mycoplasma mycoides subsp. mycoidesRecommended by Pablo Zunino based on reviews by 2 anonymous reviewersMycoplasma mycoides subsp. mycoides (Mmm), a pathogenic wall-less bacterium, is the etiological agent of contagious bovine pleuropneumonia (CBPP). This highly contagious respiratory disease may develop in severe pneumonia, with associated high mortality rates in cattle. Mmm can display different immune evasion mechanisms; in addition, a host uncontrolled inflammatory response stands for lung lesions and chronic carrier animals. Macrophages are among the most important lines of defense against Mmm of the lower respiratory tract. Although their importance in defense and immune response modulation is known, results about their role and mechanisms of action are scarce and sometimes conflicting. In the present study, Totté et al. (1) aimed to investigate the interaction of bovine macrophages (isolated from cattle peripheral blood mononuclear cells) with Mmm, under in vitro conditions. The authors highlight that the study was performed under physiological conditions (in the presence of complement prepared from the same cell donor). In their study, using different approaches, the authors provide interesting and original results, proposing a pivotal role of complement in controlling the inflammatory response, which is crucial in the CBPP pathogenesis. The authors reported that macrophages did not kill Mmm in the presence of a non-bactericidal concentration of bovine serum. However, Mmm inactivation was observed when antiserum from CBPP convalescent animals was used. They also observed that Mmm induced the production of TNF by macrophages (when a high MOI was assessed). However, complement could even abolish Mmm-induced TNF response when used at bactericidal activity concentrations. This role of complement could be combined with the development of potentially protective antibodies against particular Mmm antigens involved in the interaction with identified macrophage receptors to propose control strategies against CBPP. Overall, the study by Totté et al. provides new fundamental insight for the research on preventive or therapeutic strategies for a poorly understood disease that still represents a serious concern for livestock production. REFERENCES 1. Totté, P., Bonnefois, T., Manso-Silván, L. Interactions between Mycoplasma mycoides subsp. mycoides and bovine macrophages under physiological conditions. bioRxiv 2022.12.06.519279, ver. 2 peer-reviewed and recommended by Peer Community In Microbiology. https://doi.org/10.1101/2022.12.06.519279 | Interactions between *Mycoplasma mycoides* subsp. *mycoides* and bovine macrophages under physiological conditions | Philippe Totté, Tiffany Bonnefois, Lucia Manso-Silvan | <p style="text-align: justify;">Abstract</p> <p>We investigated the interactions of unopsonized and opsonized *Mycoplasma mycoides* subsp. *mycoides* (Mmm) with bovine macrophages *in vitro*. Mmm survived and proliferated extracellularly on bovin... |  | Microbe-microbe and microbe-host interactions | Pablo Zunino | Anonymous, Anonymous | 2022-12-09 15:12:53 | View |
MANAGING BOARD
Roey Angel
Anne Daebeler
Craig W. Herbold
Cédric Hubas
Melina Kerou
Katharina Kitzinger
David K. Ngugi










