

Improved characterization of food microbial ecosystems, especially those fermented is key to the development of food sustainability. Short-read metabarcoding is one of the most popular ways to study microbial communities. However, this approach remains complex because of the locks and biases it may entail particularly when applied to fungal communities.
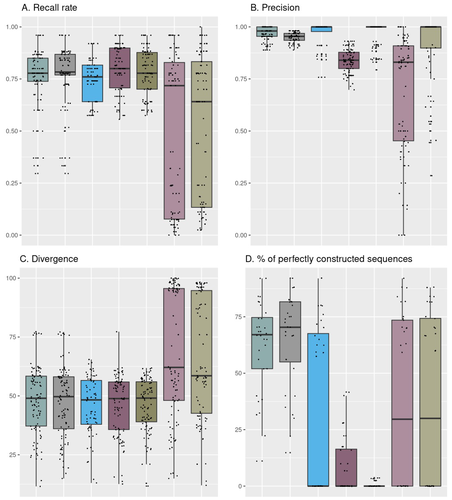
Building and using four mock communities from fermented food (bread, wine, cheese, fermented meat), Rué et al., 2023 demonstrate that combined DADA2 denoising algorithm followed to the FROGS tools gives a more accurate description of fungal communities compared to several commonly used bioinformatic workflows, dealing with all amplicon lengths. Moreover, Rué et al., 2023 provide guidance on which barcode to use (ITS1, ITS2, D1/D2 and RPB2), depending on the fermented food studied.
Practices in metabarcoding of fungi have been recently reviewed by Tedersoo et al., 2022 and their synthesis comes to the same conclusion as Rué et al., 2023. As the reference databases are far from being complete notably for food ecosystems, the development of specific sequences public databases will enable the scientific community to lift the veil on this whole area of microbial ecology.
The study conducted by Rué et al. (2023) provides a particularly detailed approach from a technical point of view, which contributes to improving the general practices in the metabarcoding of fungi. The design and the use of mock communities to compare the performances of the different pipelines is a strong point of this study. Another key element is the creation and use of an in-house database of fungal barcode sequences which improved the species-level affiliations
However, the study of fungal communities by metabarcoding is still a promising avenue of research in agri-food sciences. Thus, short-read sequencing, combined with suitable pipelines and databases, should remain of interest to the microbial ecology community (Pauvert et al., 2019; Furneaux et al., 2021).
References
Furneaux, B., Bahram, M., Rosling, A., Yorou, N. S., & Ryberg, M. (2021). Long‐and short‐read metabarcoding technologies reveal similar spatiotemporal structures in fungal communities. Molecular Ecology Resources, 21(6), 1833-1849. https://doi.org/10.1111/1755-0998.13387
Pauvert, C., Buée, M., Laval, V., Edel-Hermann, V., Fauchery, L., Gautier, A., ... & Vacher, C. (2019). Bioinformatics matters: The accuracy of plant and soil fungal community data is highly dependent on the metabarcoding pipeline. Fungal Ecology, 41, 23-33. https://doi.org/10.1016/j.funeco.2019.03.005
Rué, O., Coton, M., Dugat-Bony, E., Howell, K., Irlinger, F., Legras, J. L., ... & Sicard, D. (2023). Comparison of metabarcoding taxonomic markers to describe fungal communities in fermented foods. BioRxiv, 2023-0113.523754, ver.3 peer-reviewed and recommended by Peer Community in Microbiology. https://doi.org/10.1101/2023.01.13.523754
Tedersoo, L., Bahram, M., Zinger, L., Nilsson, R. H., Kennedy, P. G., Yang, T., ... & Mikryukov, V. (2022). Best practices in metabarcoding of fungi: From experimental design to results. Molecular ecology, 31(10), 2769-2795. https://doi.org/10.1111/mec.16460
DOI or URL of the preprint: https://doi.org/10.1101/2023.01.13.523754
Version of the preprint: 2
Dear recommender,
We have answered all reviewer comments and hope this version will meet PCI requirments.
My best regards
Delphine Sicard
Dear authors,
Could you consider the minor comments, in particular the one about methods (line 330 : Is it an OTU represented by a centroid, a Swarm seed or a denoised sequence variant ?) and the manuscript will be ready to be recommended.
Sincerely,
Caroline Strub
I only have a few minor comments and one issue which has been adressed before and is not resolved yet. Otherwise I would recommend this preprint for publication.
Ad Introduction:
Line 81: This sentence does not really reflect the findings of the Ihrmarks paper and contradicts the findings in the preprint which shows a high divergence rate for all pipelines.
L114: I guess "downside" not "downfall" is meant here.
L124: Building correct biological sequences is beside the point of traditional de novo clustering.
Ad Methods:
L330: I thank the authors for their answer but apparently no changes were made in this respect in the preprints methods. To be more explicit: A difference that can make a lot of difference, especially when talking about perfect matches to reference sequences, is what is compared with that reference sequence - is it an OTU represented by a centroid, a Swarm seed or a denoised sequence variant? This is not implicit for every pipeline and "following authors guidelines" is too unspecific for USEARCH and Qiime. I guess for USEARCH the authors refer to "recommended procedures" at https://drive5.com/usearch/manual/. There both, OTU clustering and denoising, are given which makes this reference ambiguous on how things have been done in the preprint. Similar is true for Qiime. It should not be necessary for the reader to screen the code in the supplementary just to get the information if the respective pipeline was using ZOTUs, ASVs or OTUs.
Ad Discussion:
L680: It is rather likely that all primers have missmatches with certain groups of fungi.
https://doi.org/10.24072/pci.microbiol.100007.rev21The amended version and the author's responses are satisfactory.
https://doi.org/10.24072/pci.microbiol.100007.rev22DOI or URL of the preprint: https://doi.org/10.1101/2023.01.13.523754
Version of the preprint: 1
Dear Authors,
I would like to apologize for the delay in the handling process of your preprint.
You present a novel approach to solve the issue concerning the length polymorphism of ITS1 and ITS2 sequences in metabarcoding of fungi.
Both reviewers and I agree this is a relevant study which requires moderate revision, following comments by the reviewers.
Sincerely,
Caroline Strub
This study entitled “Comparison of metabarcoding taxonomic markets to describe fungal communities in fermented food” can be divided into two sub-sections: First a comparison of mock communities of four common fungal identification markers (ITS1, ITS2, D1/D2, and RPB2) and seven bioinformatics workflows (using four different bioinformatics tools) including the most common approaches (OUT, ASV, ZOTUs) with a focus on fermented foods using four fermented food models (bread, wine, cheese, fermented meat).
The title reflects the content of the paper and the main results of the study are summarised in the abstract. The research question is very relevant to the field of food microbiology and microbial ecology and is well addressed, using relevant approaches and tools.
This paper provides an excellent contribution to the field of food microbiology. The authors demonstrate a thorough understanding of the subject matter and present a well-designed study that compares different metabarcoding pipelines and markers to determine fungal diversity in food ecosystems, with a focus on fermented foods.
The use of mock communities to validate the bioinformatics tools is a particularly strong aspect of this study, as it allows for rigorous testing of the pipelines in a controlled setting.
The comparison of four bioinformatics pipelines, including DADA2, QIIME, FROGS, and a combination of DADA2 and FROGS, is also noteworthy. The authors' demonstration of the superiority of the combined DADA2 and FROGS tools will be of interest to researchers in this field.
The paper highlights the importance of selecting appropriate markers, with the authors finding that ITS markers performed better than D1D2. The study provides guidance on the best markers for different food ecosystems, with ITS2 being best suited to characterize cheese, wine, and fermented meat communities, while ITS1 performs better for sourdough bread communities.
Overall, this scientific paper presents a thorough and well-executed study that makes a valuable contribution to the field of bioinformatics. The questions addressed are relevant to the field and the results will be of interest to researchers in agri-food sciences and microbial ecology, and the paper provides a framework for future research in this area.
I therefore recommend this paper for submission.
I also have some specific comments for the author to address:
- The relevance of figure 1 is questionable. It does not bring a substantial amount of information. Moreover, data in the text below figure 1 do not seem to confirm the data represented in figure 1: for example for meat, the text states that 4 species per genus were found for Yarrowia and Cladosporium and 2 species for Candida only. However, in figure 1, only a single dot can be found at 4 species/genus and 3 dots at 2 species/genera. Either there’s a consistency problem between the figure and the text (especially for meat) or the data are confusingly expressed.
Provide a detailed, objective report on the merits of the preprint.
Identify flaws (if any) in the design of the research, and in the analysis and interpretation of results.
Expose your concerns (if any) about ethics or scientific misconduct.
State the preprint’s strengths as well as its weaknesses. Try to consider both the technical merit and the scientific significance.
If there is something critically missing, report it.
Provide specific suggestions for improvements.